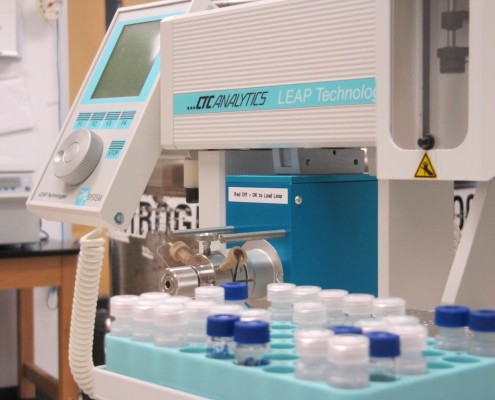
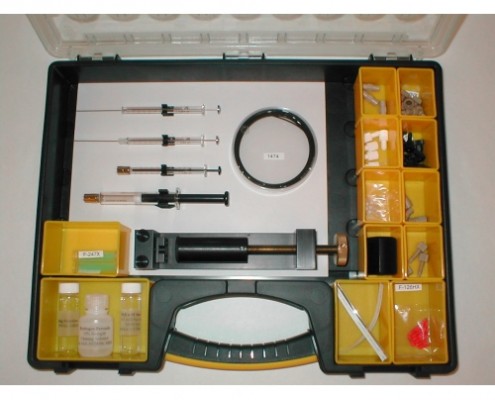
Nuclear Magnetic Resonance Spectroscopy is an important technique for the analysis of biological fluids in life science applications such as metabonomics, metabolite identification, biomarker analysis etc. Flow-NMR techniques are ideal in these applications for high-throughput screening and hyphenation with other analytical methodologies such as chromatography and mass spectrometry. Enhanced mass sensitivity, superior fluidic performance, reduced cost, and greater speed are possible using capillary-scale, microflow NMR due to the reduced sample and solvent volumes required, enabling analysis of fluids from smaller animals or any fluid that is only available in small volumes.
Featured Chapter
Chapter 16. NMR-Based Metabolomics for Biomarker Discovery
Biomarker Methods in Drug Discovery and Development, Edited by Feng Wang, Humana Press
Narasimhamurthy Shanaiah PhD, Shucha Zhang MS, M. Aruni Desilva MS, Daniel Raftery PhD
Summary
Metabolomics provides a powerful set of tools for pharmaceutical and clinical research in a number of important areas that include drug development, early disease detection, patient stratification for treatment, and information on disease processes. With its ability to discover new metabolic markers, metabolomics (as well as metabolic profiling and metabonomics) is highly effective for drug development by providing early preclinical indications of efficacy and toxicity. The most information-rich techniques currently employed in metabolomics-based studies today are nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry (MS). NMR spectra of untreated biosamples provide an overview of all metabolites present, and the complete spectrum can be used as a fingerprint of metabolic status. Analysis by multivariate statistical methods is used to identify potential biomarkers of altered metabolism that can improve the understanding of the health and disease processes. Current trends and recent advances in NMR-based metabolomics are focused on the development of advanced NMR methods, improved multivariate statistical data analysis, and a number of efforts to identify altered metabolites and pathways. Applications in the areas of toxicology, inborn errors of metabolism, cardiovascular disease, and cancer detection are described, and the prospects and the future directions of the technology are highlighted.
Excerpt
The main alternative to MS-based approaches to metabolic profiling is provided by NMR. NMR spectroscopy is especially suitable for metabolomics as it requires little or no sample preparation; is rapid, nondestructive, and noninvasive; and provides highly reproducible results. These features can provide important advantages in a number of situations. NMR spectroscopy typically have lower sensitivity compared with MS methodologies. However, with the introduction of higher field magnets (up to 900 MHz), cryogenically cooled probes (that reduce thermal noise), and microprobes equipped to handle very small (high nanoliter to low microliter) samples, methodologies that couple NMR to liquid chromatography and solid-phase extraction (19,20), sensitivity and resolution are being improved. More recently, high-resolution magic-angle spinning NMR techniques have created opportunities for the application of metabolomics to intact tissue samples (21).
Typically, 1H NMR spectra of biofluids such as urine and plasma contain thousands of signals arising from hundreds of endogenous molecules representing many biochemical pathways. Conventional measurements of the major NMR signals can be used to detect biochemical changes, but the complexity of the spectra and the presence of natural biological variation across a set of samples make the use of data reduction and pattern recognition techniques highly advantageous. The use of statistical tools such as principal component analysis (PCA), hierarchical cluster analysis (HCA), stepwise linear regression (SLR), and partial least squares (PLS) in the interpretation of metabolic profiling data set is now widespread (7,22). These methodologies are extremely helpful tools for filtering the large amounts of data and for accessing the often-subtle biochemical perturbations latent in the spectra. In addition, these approaches are used to extract single biomarkers or sets of biomarkers with the best properties for the prediction of diseases, organ function, or drug toxicity and efficacy.
Featured Speaker
Dr. Tom O’Connell, Hamner-UNC-Metabalomics Laboratory
Our Customer Partner
CapNMR + One-Minute NMR